Franz-Ulrich Hartl
Max Planck Institute of Biochemistry

Arthur L. Horwich
Yale University School of Medicine
For discoveries concerning the cell’s protein-folding machinery, exemplified by cage-like structures that convert newly made proteins into their biologically active forms.
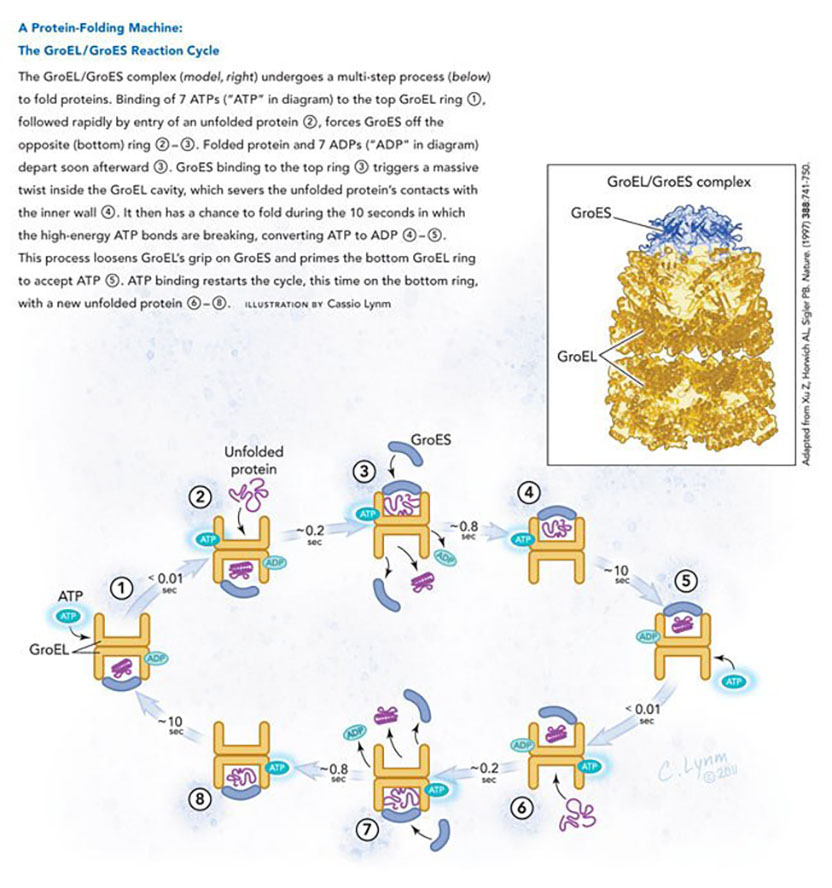
The 2011 Albert Lasker Basic Medical Research Award honors two scientists for their discoveries concerning the cell’s protein-folding machinery, exemplified by cage-like structures that convert newly made proteins into their biologically active forms. With this work, Franz-Ulrich Hartl (Max Planck Institute of Biochemistry, Martinsried) and Arthur L. Horwich (Yale University School of Medicine) toppled traditional notions of how proteins fold inside cells and established new principles that operate from microbes to humans. This previously unexplored realm holds enormous importance for basic biology and biomedicine.
Protein folding is a vital process, as it converts linear amino acid chains into the three-dimensional forms that bestow the molecules’ unique activities. Greasy regions of new proteins, however, can grab one another and create useless globs. As proteins take shape, they bury these hydrophobic parts and expose hydrophilic, or water-loving, areas. Horwich and Hartl discovered that a special apparatus encases an unfolded protein and spurs folding by harnessing the energy of ATP, the small molecule that drives reactions inside cells.
Award presentation by Titia de Lange
The 19th-century philosopher Schoppenhauer said: “Talent hits a target no one else can hit; genius hits a target no one else can see.”
The target hit by the two basic scientists we honor today concerns the question of how proteins achieve their active state in the cell. At the time of their discoveries, Ulrich Hartl and Arthur Horwich were the only ones to see that this problem was unresolved. The rest of us were blinded by the false tenet that this issue had long been settled.
The misconception about how proteins fold into their native structure originated from key observations made some 30 years earlier, by Christian Anfinsen at NIH. Anfinsen, who shared the 1972 Nobel Prize in Chemistry for his discovery, had taken a purified active enzyme and denatured it so that its long chain of amino acids would unfold. He then diluted out the denaturant and discovered that the protein could spontaneously regain its enzymatic activity. The conclusion was clear: the amino acid sequence of the enzyme’s polypeptide chain is sufficient to specify its enzymatic activity, and since the enzymatic activity depends on the three dimensional structure of the protein, it followed that a chain of amino acids can fold correctly without addition of other proteins or ATP, nature’s energy currency.
Acceptance remarks

Acceptance remarks, 2011 Lasker Awards Ceremony
Receiving the Lasker Basic Medical Research Award is a tremendous honor and a great privilege. I am pleased to receive this award together with Art Horwich, with whom I share exciting memories of the early phase of discovery in the molecular chaperone field. I am very grateful to the Lasker Foundation and especially to the Lasker Jury for selecting us, and I wish to express my gratitude to the many people who have supported and ‘chaperoned’ me along the way, especially my wife and long-term collaborator Manajit.
To be recognized by a Lasker Award is of course the dream of every biochemist or molecular biologist. I am amazed that this dream has become a reality for me. But we should not forget that our true reward as scientists is the privilege of doing something that we enjoy so deeply and that doesn’t feel like work at all. I am well aware that our discoveries were only possible because many others who came before us laid the foundation. This, and a large quantity of good fortune, allowed us to make the surprising finding, in the late 1980s, that proteins, the molecules that drive almost every biological process, reach their proper shape (their ‘fold’) by the help of other proteins, which we call molecular chaperones. This was unexpected in light of the widely held view at the time that protein folding occurs spontaneously. It was my mentor Walter Neupert at Munich University who introduced me to Art. Walter supported us with his insight and provided the environment in which our collaboration could be successful. The mechanisms we eventually uncovered are beautiful in their simplicity: A large barrel-shaped protein complex, called chaperonin, functions like a mini-test tube in the cell, allowing immature protein molecules to fold, one at a time, while being shielded from the densely populated cellular milieu. As a result, new protein chains are protected from clumping together — just like the human chaperone prevents improper interactions between young people.
I grew up in a small village in the northern part of the Black Forest. Early on in high school, I developed a strong interest in biology and chemistry. In my second year of medical school at Heidelberg University, I was given the chance to work in the biochemistry lab. I quickly realized that this was what I wanted to do, rather than practicing medicine. Who knows how many lives I saved by that decision. In any case, I was hooked by research, despite the frustrations when experiments didn’t work. I remember vividly the Hungarian guest researcher who spent hours and hours in the cold room isolating proteins. When he emerged, apparently quite exhausted, he would invariably say “Biochemie ist kein Spaziergang” (“Biochemistry is not a walk in the park”). Owing to many fortunate circumstances, I can say that for me biochemistry became the path to a very exciting and rewarding life.
In recent years, research in a number of laboratories provided evidence for the medical relevance of molecular chaperones in preventing the formation of protein aggregates that cause presently incurable diseases like Alzheimer’s, Parkinson’s, or Huntington’s disease. Hopefully, one day it will be possible to harness the power of molecular chaperones for the treatment of these diseases. Should this happen, I can honestly say it was not planned, since for the most part I just wanted to find out a bit more about the fascinating workings of cells, without any application in mind. Unfortunately, science policy makers often do not realize the potential benefits of supporting such basic research. It is to the credit of organizations like the Lasker Foundation that the public is made aware of how scientific progress is most often achieved, namely through pure curiosity-driven research.
Finally, let me say what a great pleasure and honor it is to receive this award here in New York, the city in which my wife and I have spent some of the best years of our life and where we have so many wonderful friends.
Acceptance remarks, 2011 Lasker Awards Ceremony
I thank the Lasker Foundation and the jury for this enormous honor, which goes on beyond anything I could ever have imagined. There’s a wonderful comment of Sir Isaac Newton that I think reflects my life in science: “I know not what I may appear to the world, but to myself I seem to have been only like a boy playing on the sea-shore, and diverting myself in now and then finding a smoother pebble or a prettier shell than ordinary, whilst the great ocean of truth lay all undiscovered before me.” I feel ever so lucky that we picked up the mif4 mutant pebble that informed about what is a genuinely beautiful machine that Mother Nature uses to assist the folding of proteins in cells. I feel incredibly lucky not only to have been allowed to understand the general action of this machine but to have had the thrill to actually see it at near atomic resolution and ultimately, along with others, dissect how it works. Seeing GroEL on a graphics display for the first time in fall 1993 side by side with Zbyszek Otwinowski, who solved the structure and is here today, and the late Paul Sigler was a spiritual experience.
But I love science for all the moments between these once- or twice-in-a-lifetime moments. I love Newton’s “sea-shore” — the quiet contemplation of how something might work, the collaborative discussions with Ulrich and Ming (who is here today from Taiwan) in those early times, and with everyone in my lab every day, where I work at the bench side by side with everyone. And I love the personal dirtying of my own hands in my own experiments, where failure is the common outcome, but where I invest my being, as if a boy on the sea-shore.
We have recently directed our focus to the neurodegenerative paralyzing motor disease ALS, to a form caused by protein misfolding, and I hope that we or others are able to pick up the shell that will be the understanding of the basic mechanism by which a misfolded protein causes neurotoxicity. That might lead to a rationally designed therapy for this devastating condition. I once again thank you all for honoring me today.
Interview with Franz-Ulrich Hartl and Arthur L. Horwich
Video Credit: Susan Hadary