
Michael N. Hall
Biozentrum University of Basel
For discoveries concerning the nutrient-activated TOR proteins and their central role in the metabolic control of cell growth
The 2017 Albert Lasker Basic Medical Research Award honors a scientist who discovered the nutrient-activated TOR proteins and their central role in the metabolic control of cell growth. By showing that the TOR system adjusts cell size in response to the availability of raw materials, Michael N. Hall (Biozentrum, University of Basel) revealed an unanticipated linchpin of normal cell physiology. TOR balances constructive and destructive activities to match accumulation of cell mass with nutrient supply and other growth signals, such as hormones. Disruption of the TOR network contributes to numerous human illnesses, including diabetes and cancer, and has been implicated in a wide range of other age-related disorders.
The sTORy begins
In the late 1980s, many researchers were studying the tightly choreographed process by which cells divide. Although scientists recognized that a cell must increase in content and size as part of this process, they were not focused on that aspect. Growth was thought to be spontaneous, so if raw materials such as amino acids and fatty acids were present, the thinking went, cells would manufacture macromolecules such as proteins and lipids. In this scenario, a growth regulator was unnecessary.
At that time, Hall was studying how proteins cross the membrane barrier that separates the main body of the cell, or cytoplasm, from the nucleus, where genes reside. Toward that end, he was exploring how a certain class of drugs (called immunosuppressants) obstruct T-cell activation, the process by which these immune cells rev up and proliferate. Scientists knew that these drugs somehow block transmission of a signal that travels into the nucleus to turn on certain genes. The medicines held great interest because their ability to quash inflammation helps prevent organ rejection after a transplant.
Hall hoped to discern the drugs’ then-murky mechanisms of action by exploiting the experimental power of baker’s yeast. The wisdom of studying an immunosuppressant in this single-celled creature was questionable, given the vast evolutionary distance between humans and yeast. However, one of the drugs, rapamycin (sirolimus), had originally been identified as an anti-fungal agent, so Hall thought it might prevent yeast, too, from duplicating. The first sign that his bold strategy would flourish emerged when he showed that rapamycin blocks proliferation of yeast cells, the same effect it has on mammalian cells.
To understand how rapamycin works, Hall wanted to identify proteins with which it interacts. He sought genetic variants of yeast that could divide in the presence of the chemical. Such mutants, he reasoned, might carry alterations in genes that encode rapamycin-binding proteins. The perturbations would render the drug impotent because it would be unable to grab and inactivate its usual molecular quarry.
Two previously unknown genes emerged from this process. Hall named them TOR1 and TOR2, for target of rapamycin, when he and postdoctoral fellow Joseph Heitman published the results in 1991. Hall isolated normal versions of the genes and analyzed their sequences, which turned out to encode two closely related proteins. When he engineered yeast to lack both TOR genes, he reported in 1993, the organisms got stuck early in the cell-division cycle, as they do when treated with rapamycin. This mimicry indicated that the drug obliterates TOR activity.
By a year and a half later, four research groups had identified mammalian TOR based on its ability to bind rapamycin. Originally, three of them gave the single mammalian TOR, whose sequence resembles those of its two yeast counterparts, a different name, but now it is universally referred to as mTOR. The shared function and structure bolstered the notion that TOR would perform similar tasks across the evolutionary spectrum.
TOR’s sequence established a novel family of kinases, proteins that affix a phosphate chemical group to other molecule(s), but the potential recipients and effects of that deed were unknown, as were the protein’s triggers and restraints. Presumably TOR fostered an early step of cell division, as cells that lack TOR function arrest at that point in the process.
TOR-deficient cells, however, displayed a perplexing trait. Yeast with flaws in proteins known to promote early steps of cell division swell in size because they accumulate contents without separating into two daughters cells. Yet yeast that lack TOR activity did not grow exceptionally large. This conundrum intrigued Hall, and by unraveling it, he exposed an unexpected regulatory system that governs cell growth.
Stabilizing growth economics
In 1996, Hall reported that cells with nonfunctional TOR initiate protein synthesis feebly and generate unusually small quantities of protein. Thus, the cells fail to duplicate not because TOR does a specific job in cell division, but because protein manufacture plummets in general, including for proteins that propel the cell through the division cycle.
As Hall dug deeper, he realized that TOR-deficient cells behave as if they are starving. When nutrients are scarce, yeast ramp down their building activities. They subdue protein and most messenger RNA production, remaining alive but metabolically dormant. By all criteria tested, TOR-deprived yeast were stalling out in this limbo-like state. Hall proposed that normal, intact TOR stimulates cell growth in response to nutrient availability. Several years later, he showed that it sits at the crux of a signaling pathway that receives information about nutrient status and sparks downstream events that tailor gene activity in response.
Hall and others began untangling how TOR tunes protein synthesis and other activities. An explosion of work has established that TOR—in yeast and mammals—encourages macromolecule production in many ways and on multiple levels. In favorable circumstances, it not only incites protein and nucleic acid synthesis, but promotes manufacture of their subunits and assembly of machines such as ribosomes that make them. It also fosters synthesis of lipids, key ingredients of the cell membrane, which must expand as cells bulk up. Conversely, TOR dampens processes that break apart large molecules to recycle their components.
In addition to unveiling TOR’s growth-related deeds, scientists have begun to probe mechanisms by which nutrients influence its behavior. For example, David M. Sabatini (Massachusetts Institute of Technology and Whitehead Institute for Biomedical Research) has shown how amino acids prompt mTOR activation at a particular site inside the cell.
Positioning growth
Stimulation of metabolic pathways is a key aspect of growth, but physical changes must also occur. As yeast cells head toward division, a bulge appears at a particular site on their surface. It expands and eventually pinches off to form a new daughter cell. This budding process involves reorganization of internal filaments, composed largely of the protein actin, at a specific location. By molding this framework, the cell thus shapes itself to promote reproduction.
Through numerous strategies, Hall established that TOR2 controls these actin rearrangements. The initial hint came from the observation that the two TOR genes are not interchangeable, despite significant sequence identity. Hall’s early genetic experiments showed that TOR2 could substitute for TOR1, but TOR1 could not substitute for TOR2. He proposed that two signaling pathways operate, one shared by TOR1 and TOR2, and one unique to TOR2. The first instructs the cell when to shift into a constructive metabolic mode—when nutrients are plentiful—and the second tells the cell where to grow—at the bud site. In this way, he suggested, the TOR system ensures that growth occurs only at specific times and places.
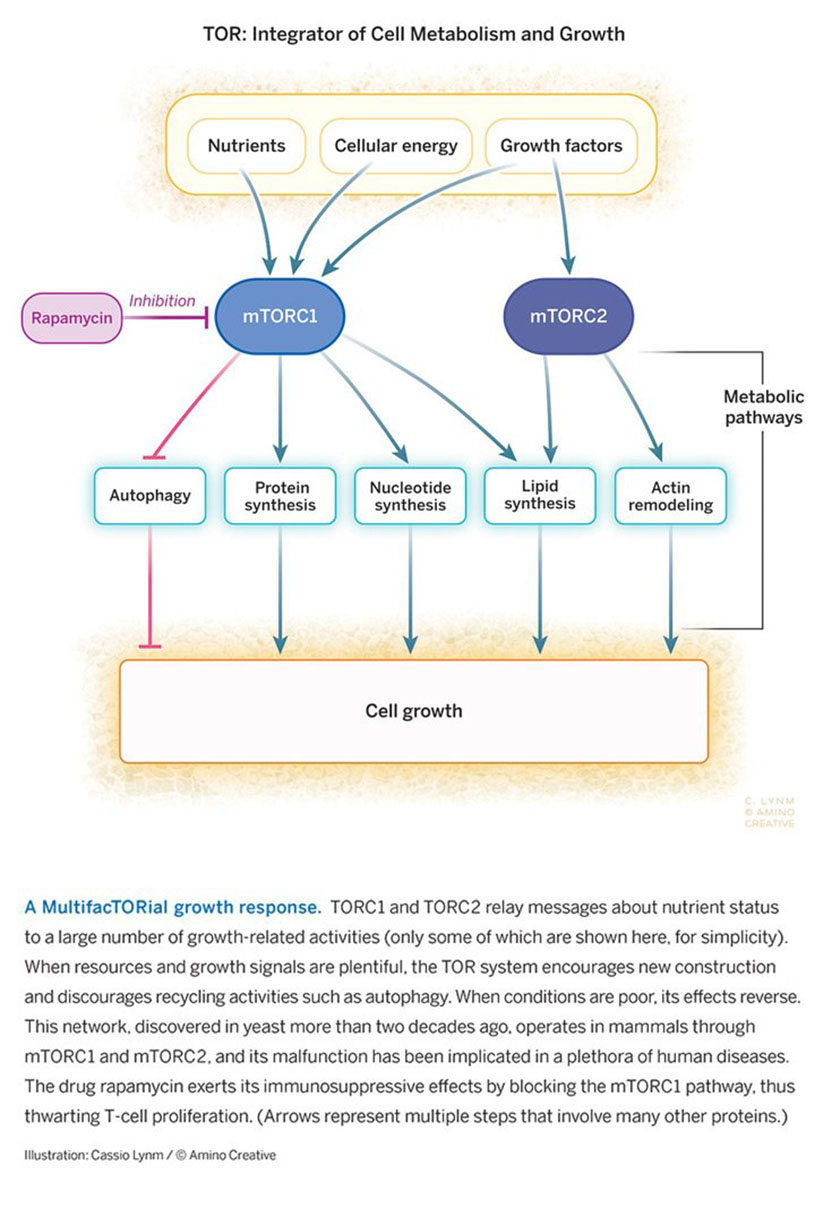
In 2002, Hall uncovered the physical basis for these pathways. He identified and characterized two molecular assemblies, distinguished by different protein constituents. The first, TOR Complex 1 (TORC1), contains TOR1 or TOR2 plus a particular combination of other proteins, and it exerts the many metabolic effects associated with TOR; the second, TOR Complex 2 (TORC2), contains TOR2 plus a different combination of proteins, and it executes the actin-associated, TOR2-unique function as well as lipid synthesis (see Figure) and other tasks that can have mechanical influence. This work revealed why TOR2 can signal through both pathways and TOR1 can signal through only one. It also explained why rapamycin foils the shared pathway but not the unique one, as the drug does not bind TORC2.
At this point, only a TORC1-type of complex had surfaced in mammalian cells, but Hall wondered whether mammalian TORC2 might have eluded detection because of how scientists had been studying TOR function—by adding rapamycin and documenting its effects. If mammalian TORC2 physiology mirrored that of yeast, he reasoned, the strategy would have left TORC2 untouched, as the complex is insensitive to the drug.
Using methods that did not involve rapamycin, Hall and others showed in 2004 that TORC2 exists in mammalian cells and controls actin remodeling. Like yeast TOR2, the single mTOR forms two distinct complexes, mTORC1 and mTORC2. Evolution has thus retained the dual signaling pathways by which TOR governs cell growth. At some point, the ancestral system picked up additional inputs—hormonal signaling from insulin, for example—and we now know that mTOR helps multicellular creatures coordinate cell growth across a whole body.
Hall discovered TOR and thus unearthed a master regulator that links resource availability with myriad aspects of metabolism and growth. These processes are essential to all living things, so it is no surprise that TOR touches a tremendous array of human pathological conditions in which some facet of them has gone awry. Scientists are putting these findings to work—developing mTOR inhibitors, for example, for certain cancers. By studying a clinically useful drug in yeast, Hall illuminated a signaling pathway with profound medical implications for humans.
by Evelyn Strauss
Key Publications of Michael N. Hall
Heitman, J., Movva, N.R., and Hall, M.N. (1991). Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast. Science. 253, 905-909.
Kunz, J., Henriquez, R., Schneider, U., Deuter-Reinhard, M., Movva, N.R., and Hall, M.N. (1993). Target of rapamycin in yeast, TOR2, is an essential phosphatidylinositol kinase homolog required for G1 progression. Cell. 73, 585-596.
Barbet, N.C., Schneider, U., Helliwell, S.B., Stansfield, I., Tuite, M.F., and Hall, M.N. (1996). TOR controls translation initiation and early G1 progression in yeast. Mol. Biol. Cell. 7, 25-42.
Beck, T., and Hall, M.N. (1999). The TOR signalling pathway controls nuclear localization of nutrient-regulated transcription factors. Nature. 402, 689-692.
Loewith, R., Jacinto, E., Wullschleger, S., Lorberg, A., Crespo, J.L., Bonenfant, D., Oppliger, W., Jenoe, P., and Hall, M.N. (2002). Two TOR complexes, only one of which is rapamycin sensitive, have distinct roles in cell growth control. Mol. Cell. 10, 457-468.
Hall, M.N. (2016). TOR and paradigm change: cell growth is controlled. Mol. Biol. Cell. 27, 2804-2806.
González, A., and Hall, M.N. (2017). Nutrient sensing and TOR signaling in yeast and mammals. EMBO J. 36, 397-408.
Award presentation by Bruce Stillman
In 1839, Charles Darwin wrote “it appears to me that nothing can be more improving to a young naturalist than a journey in a distant country”. Like the geographic travels to which Darwin was referring, it seems to me that a different journey, one of scientific discovery, can also be improving to a young scientist. Today we celebrate a talented yeast geneticist who has traveled a very interesting scientific journey.
In parallel to Darwin’s 1835 travels to the now famous Pacific Islands of Galapagos, our journey today starts with a voyage to another Pacific Island. In the mid 1960s, Canadian scientists set out on an expedition to study the health of the native population of the remote Polynesian Island of Rapa Nui, commonly known as Easter Island. Instead of collecting finches, they gathered soil samples that were later shown to contain a Streptomyces bacterium that synthesized a chemical natural product called rapamycin. Rapamycin, together with another soil-derived natural product called FK506, attracted much attention because of their potent immunosuppressant activities, enabling their use for kidney transplants, for the treatment of ulcerative colitis and treating certain cancers. But how did these drugs work?
This journey is basic science at its highest level.
Both rapamycin and FK506 were known to bind to a protein called FKBP that was suspected of being the target of these drugs. But Hall and others knew that there were inconsistencies in how FK506 was linked to rapamycin’s immunosuppressive action. Thus many groups were trying to identify the true target of rapamycin, mostly using mammalian cells.
Hall, trained as a yeast geneticist, decided to take a different path. He knew that rapamycin had anti-fungal activities and in 1991 he showed that it was toxic to baker’s yeast. With his medically trained student Joe Heitman, Hall isolated spontaneous mutants of yeast that were resistant to rapamycin. Many mutations turned out to map to the gene encoding the yeast FKBP. But rare, dominant mutations in two previously uncharacterized genes revealed themselves, leading to the cloning of TOR1 and TOR2, TOR standing for target of rapamycin. Hall surmised that TOR1 and TOR2 would be conserved, and noted that they would exist in T cells and be the target for rapamycin’s immuno suppressive activities.
Like all great discoveries, these initial results in 1991 prompted many new journeys, not just for Michael Hall, but for many other scientists. These combined scientific journeys have yielded stunning results.
At the time when Hall first identified the TOR genes, it was then assumed that cell growth was ineluctable, limited only by the available nutrients and not controlled.
In stark contrast, when the growth of yeast cells was arrested early in the cell division cycle with rapamycin or with a conditional mutation in the TOR genes, the cells appeared as if they had been starved of nutrients, even though abundant nutrients were available. Hall presented his unpublished data in a public seminar in Vienna, leading to discussions with Kim Nasmyth, a noted cell cycle expert. Prompted in part by these discussions, Hall demonstrated that the TOR proteins control a signaling pathway. This pathway regulates the translation of critical messenger RNAs that synthesize proteins essential for cell growth.
Importantly, Hall also suggested that the TOR proteins sense nutrient availability and this is how they control cell growth. In a 2016 retrospective, Hall noted that this paradigm shifting, 1996 paper was rejected no less than 7 times. It was eventually accepted in the then new journal Molecular Biology of the Cell; his paper being edited by Lee Hartwell, a Lasker winning yeast geneticist. Lee, like Michael Hall, was completely receptive to paradigm shifts, unlike the other journal reviewers and editors. Lessons from this part of the story abound, but two are obvious. First, sharing unpublished results with smart colleagues can be of great value. Second, not publishing in so-called high profile journals can still result in the highest possible recognition of one’s work.
It soon became apparent that mammalian cells do indeed have homologs of TOR1 and TOR2, now called mTORs. They are components of very large protein complexes that control the synthesis of cellular components such as proteins, lipids and nucleic acids, the core building components required for all cells to grow. They also prevent degradation of cellular components by a process called autophagy that is induced when cells are starved of nutrients and need to survive. Upstream, the mTOR complexes are responsive to many cell signals such growth factors including insulin, to the availability of essential amino acids, to the state of the cell’s energy and even to the availability of oxygen. mTORs are in essence the cell’s monitors of feast or famine, growth or no-growth.
Since the TOR journey began about a quarter of a century ago, whole body physiological roles for mTOR complexes have been uncovered. They coordinate nutrient homeostasis in the liver. They monitor the growth of muscles in response to exercise and they are involved in the accumulation of fat in adipocytes. Thus, having just finished your lunch, you should venerate your mTORs.
Michael Hall has had an interesting geographic journey, born in Puerto Rico, living as a child in Venezueala and Peru, schooling and university studies in the U.S., and eventually journying to the Biozentrum in Switzerland. It not this geographic voyage, but his scientific journey, a journey that broke fresh ground, a trek that established a new paradigm, it is this journey that we celebrate today and for which he receives the 2017 Albert Lasker Basic Medical Research Award.
Charles Darwin, himself, would have been impressed.
Acceptance remarks by Michael N. Hall
I am deeply honored and delighted to receive the Lasker Basic Medical Research Award. How did this come to be? To answer this question, I have to tell you a short story.
A few years ago, a radio journalist asked me, “What is more important, your wife or your science?” I brushed off this seemingly silly question by answering half in jest, “My wife is more important, science is my mistress.” In truth, the question was not so silly because it made me realize that my wife Sabine and Science are the two great love affairs of my life.
As an independent investigator, I re-discovered my passion for Science and offspring ensued – talented students and postdocs. It was these students and postdocs, in some cases rebellious, that came up with creative ideas such as using yeast genetics to study the mode of action of drugs developed for use in humans. This and other ideas in turn led to the discoveries that are being celebrated today.
I would like to end with some words of thanks. I acknowledge Science itself. Science has been a sanctuary – a place of serene logic and comfort – where I have found refuge in an often irrational world. Where would I be without it? More importantly, where would humanity be without science? I thank the Albert and Mary Lasker Foundation for celebrating science in general, and our science in particular. There is no greater reward for a scientist than to be recognized by one’s peers. Last but certainly not least, I thank my wife Sabine. The scientist-philosopher Jacob Bronowski wrote in Science and Human Values (1956) that, “as a profession, science attracts men whose temperament is grave, awkward, and absorbed.” Although these might be good traits for a scientist, they are not necessarily good traits for a husband. Despite my absences and shortcomings, Sabine has always been there to provide support and encouragement. I would not be here without her.
2017 Basic Award video
Video Credit: Flora Lichtman
