Israeli biochemist Avram Hershko arrived in the US in 1969, eager to begin his postdoctoral research at the University of California, San Francisco. But when he walked into the lab where he was to spend the next two years, he was taken aback. About 20 other postdocs were already there, all working on the same project to understand how cells synthesize a particular protein. “It was a bit too crowded,” he says.
Hershko wanted more scientific freedom, so he asked his supervisor whether he could work on something else. His supervisor suggested he study how cells break down proteins, a topic Hershko and most other researchers hadn’t given any thought to. “Nothing was known about it,” he recalls. But he was quickly hooked. The subject “became an idée fixe,” he says. “I wanted to figure out how this works.”
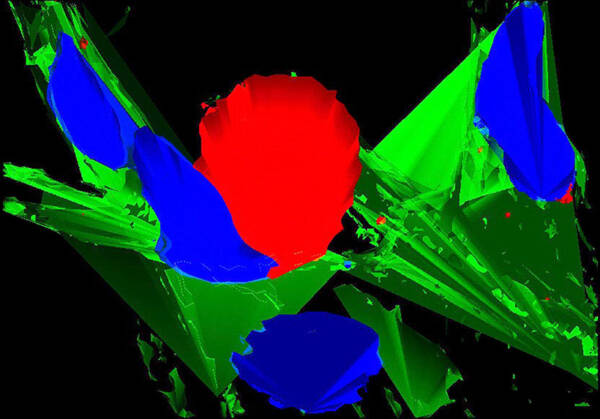

Fluorescent deconvolution micrograph of brain tissue from a patient with Parkinson’s disease. The protein ubiquitin is red, cell nuclei are blue and connective tissue is green. Ubiquitin attaches to other proteins to signal that they need to be degraded. A failure of this process is thought to contribute to the progression of Parkinson’s disease.
Courtesy of Roger J. Bick, Brian J. Poindexter, Mya C. Schiess / UT-Houston Medical School / Science Source
Hershko continued to investigate protein breakdown after he returned to Israel in 1971 and started his own lab at the Technion – Israel Institute of Technology. Graduate student Aaron Ciechanover soon joined his research group, and together they uncovered an intricate signaling network inside cells: the ubiquitin system. About 8% of human genes code for proteins that are part of that system, which is crucial for functions as diverse as cell division, defense against invading pathogens, blood vessel growth, and DNA repair. For their work, Hershko and Ciechanover earned a share of the 2000 Albert Lasker Basic Medical Research Award and the 2004 Nobel Prize in chemistry.
This is the second of two articles on Lasker winners who probed cell signaling. The seven winners profiled here revealed a key on–off switch for proteins, identified two messengers that stimulate immune cells to fight pathogens, and helped decipher some of the internal signals that enable cells to coordinate and manage their activities.
Marked for Destruction
At first, most scientists were skeptical of Hershko’s research. Their doubts only made the possibility of working with him more appealing, Ciechanover says.
“It looked to me kind of adventurous.”
The pair’s first experiment, tracing the breakdown of a specific protein in cytoplasm removed from immature blood cells, hinted at unexpected complexity in the process. Hershko and Ciechanover sorted the proteins in the cytoplasm into two groups, but neither group alone could demolish the target protein. When the pair submitted a paper describing that result to a scientific journal, a reviewer “tore it into tiny pieces,” Ciechanover says.
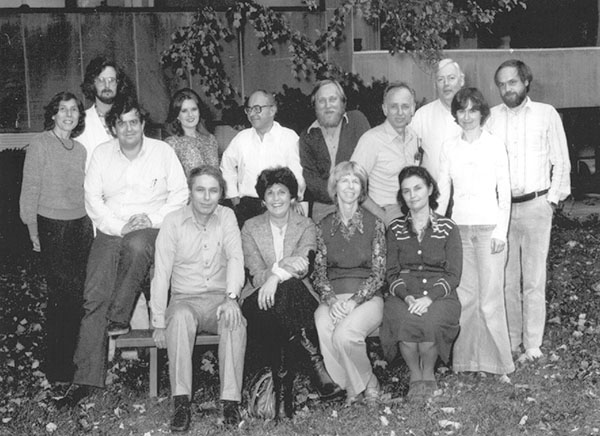

Hershko (first row, far left) and Ciechanover (second row, far left) during a sabbatical they spent at the Fox Chase Cancer Center in Philadelphia in 1979. That year they discovered that cells mark proteins for destruction by attaching ubiquitin.
Courtesy of Avram Hershko
Certain they were onto something, Hershko and Ciechanover eventually got the paper published and continued the research. Their next step involved isolating one of the process’s essential proteins: a small protein known as ubiquitin. “The purification of ubiquitin was the major breakthrough,” Ciechanover says. Once he and Hershko nabbed it, they could unravel how it functions. They learned that ubiquitin latches onto certain proteins and serves as a signal for the cell to destroy them. But ubiquitin requires three accomplice proteins to work, they found: E1 activates ubiquitin, readying it to glom onto a doomed protein. E2 serves as an escort and delivers ubiquitin to E3, which attaches it to the target. Hershko and Ciechanover performed their experiments mainly in the test tube.

Varshavsky at the 2000 Lasker Awards
The third researcher who shared the 2000 Lasker Award with them, Alex Varshavsky, now at the California Institute of Technology, filled a critical gap by extending the study of ubiquitin to mammal and yeast cells.
The ubiquitin system is indispensable for cells, enabling them to cull worn-out or incorrectly folded proteins. It also gives cells precise control over certain processes. For instance, a protein called cyclin B stimulates cells to begin mitosis, in which they parcel out their chromosomes and then divide. Once mitosis is almost complete, however, cells need to get rid of cyclin B. As Hershko and colleagues found, cells tag cyclin B with ubiquitin during mitosis, ensuring that it gets removed at the right time.

Avram Hershko (right) and Aaron Ciechanover pore over the results of an experiment circa 2000.
Courtesy of Avram Hershko
Scientists have since discovered that ubiquitin molecules don’t always mark a protein for destruction. They also can steer the protein to a different cellular location, alter its function, and affect how it interacts with other proteins. Ubiquitin is the basis for a more complex cellular language, which researchers term the ubiquitin code.
Proteins’ On–Off Switch
Ubiquitin isn’t the only molecule that cells can stick onto proteins to alter what they do. Two of the researchers who shared the 1989 Albert Lasker Basic Medical Research Award, Edwin Krebs and Yasutomi Nishizuka, helped reveal a cell signaling mechanism that relies on the molecule phosphate.
Edwin Krebs, who worked at the University of Washington for most of his career, said people often mistook him for the famous biochemist and Nobel laureate Hans Krebs. Edwin lived up to the name, performing Nobel-winning work of his own that revealed cells modify proteins’ functions by adding phosphate groups and by stripping them off—opposing processes known as phosphorylation and dephosphorylation, respectively.

Edwin Krebs (left) and his colleague Edmond Fischer, shown here in 1992 after they won the Nobel prize, surprised scientists by discovering that adding phosphates to proteins can alter their activity.
Courtesy of University of Washington
Krebs and his longtime collaborator, Edmond Fischer, made the discovery while investigating a muscle enzyme that came in two varieties, one of which was inactive. In 1955, the scientists determined that cells could switch the enzyme on by affixing a phosphate group and switch it off by removing the phosphate. The discovery was a surprise, Fischer later wrote, because
“nobody could imagine that the phosphorylation of an enzyme could be involved in its regulation.”
The discovery was a landmark because, as other researchers have discovered, phosphorylation is ubiquitous during cell signaling. When the hormone insulin latches onto a receptor on the surface of a cell, for instance, it sets off a cascade of phosphorylation reactions.

Nishizuka discovered the protein kinase C family of enzymes that phosphorylate other proteins to control their functions.
Courtesy of National Cancer Institute
Building on Krebs and Fischer’s discovery, Yasutomi Nishizuka, at Kobe University in Japan, unearthed the protein kinase C family of enzymes, which phosphorylate an array of proteins during cell signaling and help orchestrate functions such as cell division, muscle contraction, blood clotting, and inflammation.
Nishizuka and colleagues first isolated the proteins in the late 1970s, but their 1982 paper on the connection between protein kinase C and cancer “catapulted [the molecule] to the forefront of signaling,” as one researcher wrote. The scientists showed that certain cancer-causing molecules switch on protein kinase C enzymes. Researchers now know that some protein kinase C varieties promote cancer, but other types inhibit tumors. The proteins may play a role in other diseases, including diabetes, atherosclerosis, and Alzheimer’s.
Get Me a T Cell, Stat!
All signaling systems depend on specificity—a cell has to perform the correct actions once it receives a message. How immune cells such as B cells and T cells, which communicate by using chemical messengers called cytokines, generate the right responses was a mystery until the work of the 2002 Lasker Special Achievement Award winner James Darnell Jr., a biologist at Rockefeller University in New York City. He discovered two signaling intermediaries, STAT1 and STAT2, that enable cells to react to cytokines and other signaling molecules. And if that weren’t enough, Darnell also performed pioneering work on RNA. His dual research breakthroughs and his fostering the careers of many young scientists explain why he won the Special Achievement Award.
The research that led to the discovery of the STAT proteins started when Darnell and colleagues asked how cells turn on specific genes. For example, liver cells switch on different genes—and thus produce different messenger RNAs, the molecules that contain the instructions for synthesizing proteins—than brain and spleen cells. To determine what controls activation of those genes, the researchers dosed liver cells with interferons, alarm cytokines released by cells invaded by viruses. Interferons spurred the cells to pump out specific messenger RNAs, Darnell and his team found.

James Darnell is in his lab at Rockefeller University in 1989.
Courtesy of John Sholtis
They took eight years to figure out how. They eventually determined that interferons stimulate cells to phosphorylate the two STAT proteins, which then move into the nucleus. There the proteins latch onto DNA sequences near specific genes and turn those genes on. Darnell said his wife inspired the name for those fast-acting proteins because “stat” is hospital lingo for doing something quickly.
Researchers now know that seven types of STAT proteins exist and that more than 50 kinds of cytokines signal through them, helping coordinate processes such as regulating inflammation, producing new white blood cells, and responding to pathogens. Messenger molecules that control other functions such as growth and forming new blood vessels also act through STATs. Faulty signaling by STATs and the proteins they interact with drives illnesses such as inflammatory bowel disease and some cancers.
Clue from the Ice
A cell in your body has a lot of decisions to make: whether to grow or divide, how fast to set its metabolism, and even whether to live or die. Michael Hall of the University of Basel in Switzerland received the 2017 Albert Lasker Basic Medical Research Award for identifying a molecular impresario known as target of rapamycin (TOR) that helps cells make such decisions. Like many other important proteins, TOR has a dark side as well. It goes awry in illnesses such as cancer, diabetes, and Alzheimer’s disease, and it may help drive aging.
Hall uncovered TOR’s central role thanks to some yeast cultures languishing in his freezer. In late 1990, a postdoc in Hall’s lab was leaving for a new job, so he stashed the mutant yeast cultures he had been working on in the freezer. Hall had no plans to study the cultures further, and “they could have been forgotten,” he says. Only after another postdoc’s project failed did Hall and his colleagues remember the yeast and dig the cultures out.
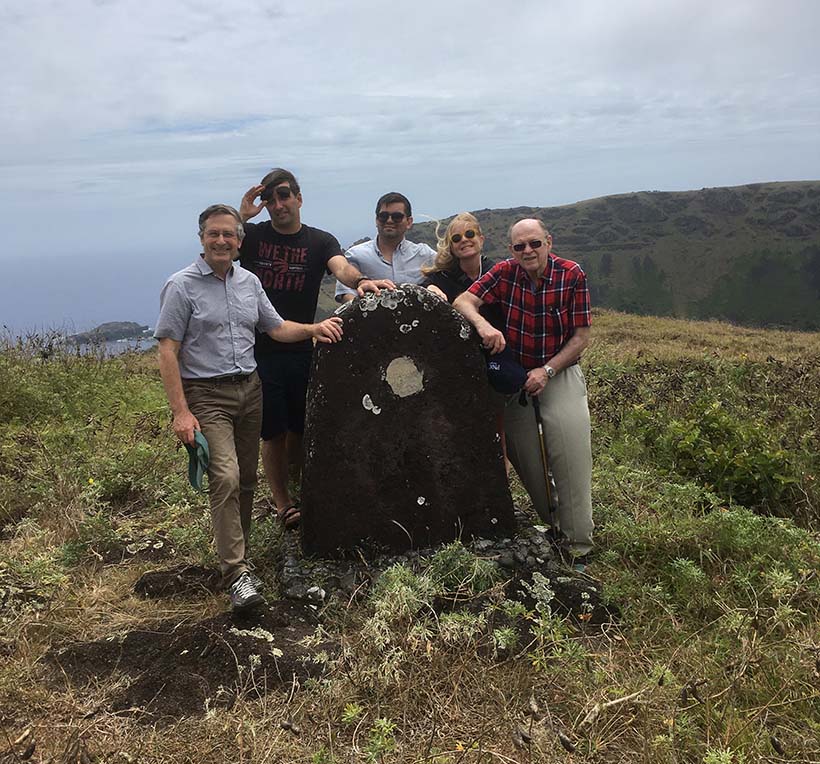

Rapamycin, a drug that targets TOR and helps prevent rejection of transplanted organs, comes from soil bacteria originally collected on the island of Rapa Nui in the South Pacific. This 2019 photo shows Michael Hall (far left) and colleagues at the site where scientists dug up the first samples of the bacteria.
Courtesy of Sabine Hall
Before the yeast cultures had gone into the deep freeze, Hall and his team had used them to investigate rapamycin, then a promising new drug derived from a bacterium native to Easter Island in the Pacific Ocean. Yeast dosed with the drug often stop dividing. The researchers identified yeast strains that shrugged off rapamycin, finding that they carried mutations in two genes, which the scientists dubbed TOR1 and TOR2. That result suggested that rapamycin works by blocking versions of TOR.
After reviving the mutant yeast cultures, Hall and his team isolated the genes in yeast for the two TOR proteins (humans and other mammals have only one). But they still didn’t know what TOR did. Hall suspected that the protein managed cell division, but when he proposed the idea to colleagues at the University of Vienna, they savaged it.
“I let them pummel me,”
he says, and after wrangling for hours the scientists came up with an explanation that better fit the data: that TOR governs cell growth.
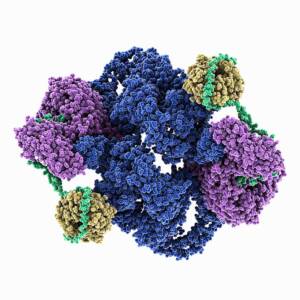
Molecular model of the structure of human mTOR Complex 2. Shown are the serine/threonine-protein kinase mTOR (blue), the target of rapamycin complex subunit LST8 (yellow), the rapamycin-insensitive companion of mTOR (purple) and the target of rapamycin complex 2 subunit MAPKAP1 (green).
Courtesy of LAGUNA DESIGN / Science Source
The hypothesis was controversial, however, because at the time scientists thought that cell growth had no control switch, Hall says. Instead, researchers assumed that cells haphazardly accumulated the proteins and other molecules they needed to get bigger. Hall and his team disproved the conventional wisdom. TOR carefully adjusts processes such as protein synthesis so that cells have the wherewithal to enlarge. “Cells have to know when to grow and how big to grow,” Hall says. TOR allows them to make those choices. But if Hall hadn’t remembered those yeast cultures sitting in his freezer, the research that led to this discovery “could have died out,” he says.
By tapping into cellular communications, the seven researchers profiled here opened up new treatments for cancer and other diseases. For instance, patients with the blood cancer multiple myeloma can benefit from three drugs that prevent destruction of ubiquitin-tagged proteins, and at least 10 other compounds designed to disrupt the ubiquitin pathway are in clinical trials. The discovery of phosphorylation had an even bigger medical impact. Now 120 approved drugs target the process, including the well-known leukemia treatment imatinib (Gleevec), as well as compounds for fighting breast, lung, colon, and kidney cancers. And researchers may be able to develop even more therapies if they listen carefully to the messages cells are sending.
By Mitchell Leslie



